Membrane protein characterization in microvesicles
Context: MPs are encoded by one third of human genes1 and play crucial roles in many biological processes (e.g. transport of ions and molecules, control of transmembrane potential, energy generation and transduction, signal recognition and transduction, catalysis of chemical reactions). Therefore, they are the major targets for validated drugs, current and predictable drug discovery2. There is an increased interest in their in vitro study but their functional and structural studies are limited due to their physicochemical properties (hydrophobicity), their low abundance in native membranes, and the difficulties encountered to express them in classical systems (toxicity, low expression levels, misfolding) and reconstitute them into proteoliposomes.
Analytical solutions, limitations and current challenges: A variety of expression systems have been developed to gain sufficient functionally active material but they have their respective advantages and weaknesses3. Among them, the bacterial expression systems represent the first trials because they are easier to handle, inexpensive, give higher yields and a facilitated scale-up to higher volume production. Two complementary systems can be tested in parallel: the oldest and most exploited system Escherichia coli, and Lactococcus lactis; some studies already combined them with functional characterization in L. lactis and structural analysis with E. coli4. L. lactis has emerged in the last decade and proved to be a good alternative to E. coli5. Around one hundred MPs from diverse origins, topologies and functions have been expressed in L. lactis using the NICE (NIsin Controlled gene Expression) system and among them, some human proteins6,7. Three 3D structures of prokaryotic MPs (up to 53 kDa and 11 α-helices) have been recently published. L. lactis presents several advantages over E. coli: i) a facilitated expression of (difficult) MPs; ii) the absence of inclusion bodies formation; iii) only 1 glycolipid-rich membrane; iv) no endotoxins and low protease activity; and, v) more interestingly, the lactic acid bacterium is food grade (used in fermented food production), i.e. generally regarded as safe 8-9(GRAS).
Today, depending on the nature of the protein, diverse functional assays based on chemical, biochemical and biophysical techniques can be performed on various biological materials such as intact cells, membrane fractions, proteoliposomes (after solubilization with detergents and purification steps), purified MPs. Nevertheless, most of them require labelling, large amounts of proteins and/or are performed on rudimentary reconstitutions of MPs in lipidic environment. Developing new label free biophysical technologies compatible with lipid presence in the sample for functional studies and small quantities of proteins is still a challenge for people working with membrane proteins.
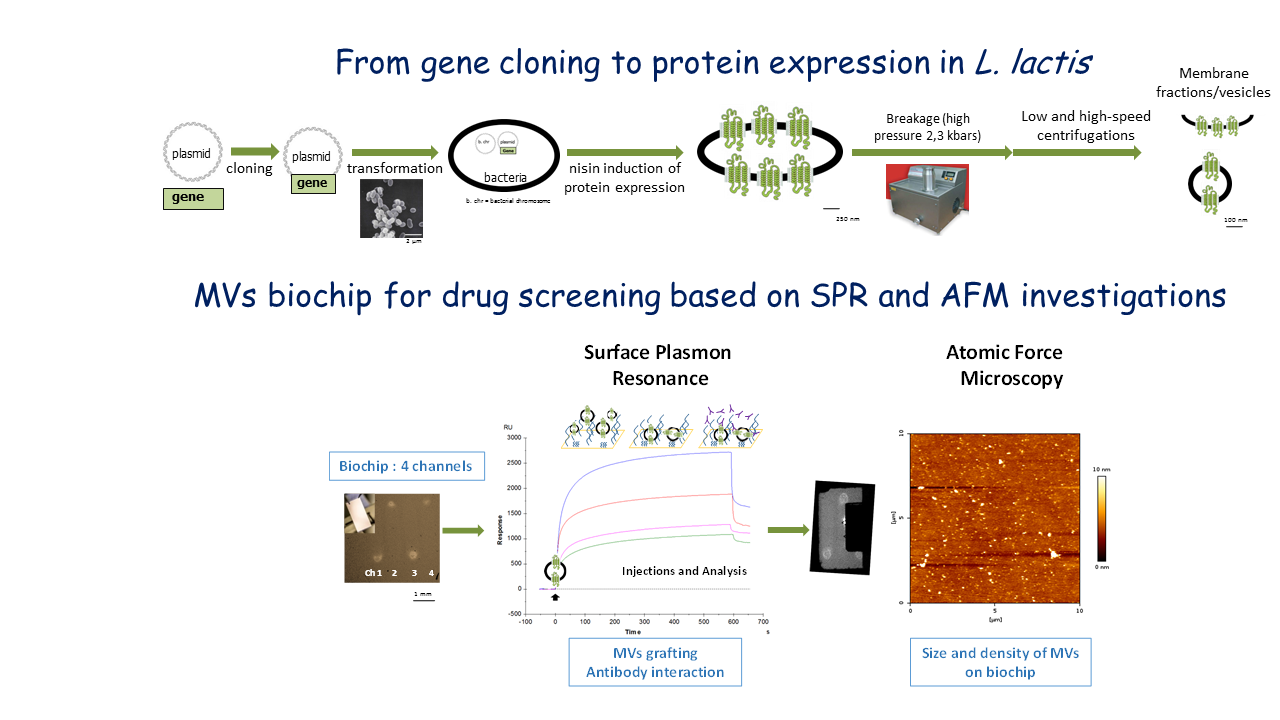
Contribution of our group to this field: A label-free NBA (nanobiocharacterization) platform of blood microparticles has been recently developed in our group based on the combination of Surface Plasmon Resonance (SPR) analysis, Atomic Force Microscopy (AFM)10 and mass spectrometry (MS) approaches. SPR detect and quantify molecular interactions in real-time on a surface, even in complex media11-12. Atomic force microscopy allows deep characterizations in size and shape13-15. Moreover, MS engaged on the chip after selective capture on arrays is the way to reach sample protein signatures in different conditions (proteomic profiling, cell signaling pathway…). This coupling “SPR/AFM/MS” allows to correlate the average measurement at the macroscale of biological activity on the chip and the in situ visualization of nanoscale biological events and protein signatures. This platform will be adapted to MP characterization in bacterial microvesicles and complete from a nanophysical point of view the panel of available techniques for their functional studies in lipidic environment.
Keywords: Surface Plasmon Resonance, Atomic Force Microscopy, Mass Spectrometry
Financial and technical supports: Région Franche-Comté (Project Emergence/Excellence) (2016), MIMENTO and CLIPP Platforms.
Contacts: A. Frelet-Barrand, C. Elie-Caille, W. Boireau
References:
1. Wallin E. and Von Heijne G (1998) Protein Sci. 7:1029-38.
2. Rucevic M. et al. (2011) Electrophoresis 32:1549-64.
3. Junge F. et al. (2008) Cell Mol Life Sci. 65:1729-55.
4. Seeger M.A. et al. (2012) PLoS One 7:e37845.
5. Bernaudat F., Frelet-Barrand A et al. (2011) PLoS One 6: e29191
6. Bakari S., …Frelet-Barrand A (2014) Chapter 5 “Membrane Proteins Production for Structural Analysis” (ed. I. Mus-Vuteau; Springer eBook) p107-132.
7. Bakari S, …. Frelet-Barrand A (2016). Mol Biotechnol. 58:299-310. doi: 10.1007/s12033-016-9928-z.
8. Kunji E.R. et al. (2003) Biochim Biophys Acta 1610:97-108
9. Mierau I. et al. (2005) Microb Cell Fact. 4:16.
10. Obeid S. et al. (2016) Bioelectron. pii: S0956-5663(16)30856-9. doi: 10.1016/j.bios.2016.08.100.
11. Remy-Martin F. et al, (2012) Anal. Bioanal. Chem. 404(2), 423-432
12. Rouleau A. et al (2012), Sensors, 12(11), 15119-1513
13. Ewald M. et al (2015), Nanotechnology, 25(29) 295101
14. Heu C. et al (2012), J. Struct Biol, 178(1), 1-7
15. Berthier A. et al (2011) J. Molec. Recogn. 24(3),429-435

